Team 7b - Cell Plasticity in Development and Cancer (DevCan)
Team leaders : Vincent GUEN (INSERM), Eloïse GRASSET (CNRS)
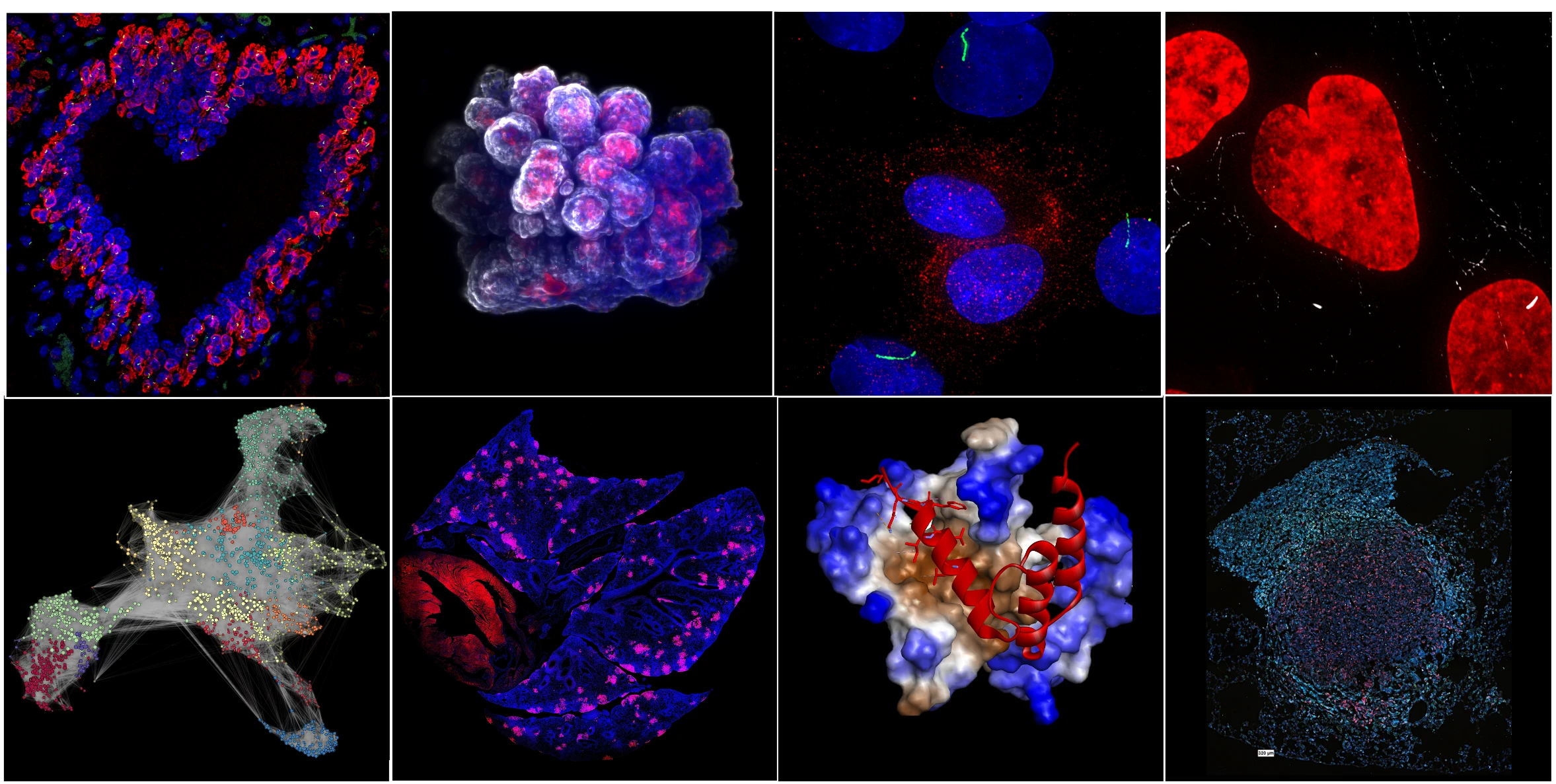
The laboratory is interested in understanding mechanisms of cell plasticity in development and disease. We leverage and develop cutting-edge bio-imaging methods and single-cell technologies together with innovative animal models, organoids, and chemical tools to map and understand cell state transitions including in ciliopathies and cancers.
Over the past decade, our research has focused on phenotypic plasticity of cells driven by epithelial-mesenchymal transition (EMT) and ciliary signaling. EMT is a cell-biological program that enables epithelial cells to acquire an array of mesenchymal-like phenotypes. The transitional phenotypic states arrayed along the E-to-M axis are frequently referred to as "EMT states". Cells residing at intermediate positions along the E-to-M spectrum of phenotypes are known as hybrid epithelial-mesenchymal (E/M) states.
Our research established that hybrid E/M states represent cells that are poised to enter into the stem cell state through formation of primary cilia, a signaling center assembled at the cell surface. We demonstrated that EMT-induced ciliary signaling can contribute to normal developmental processes, as well as to formation and therapeutic resistance of aggressive forms of carcinomas. We also established that hybrid E/M states represent cells in a high plasticity state that fuel cancer metastatic colonization.
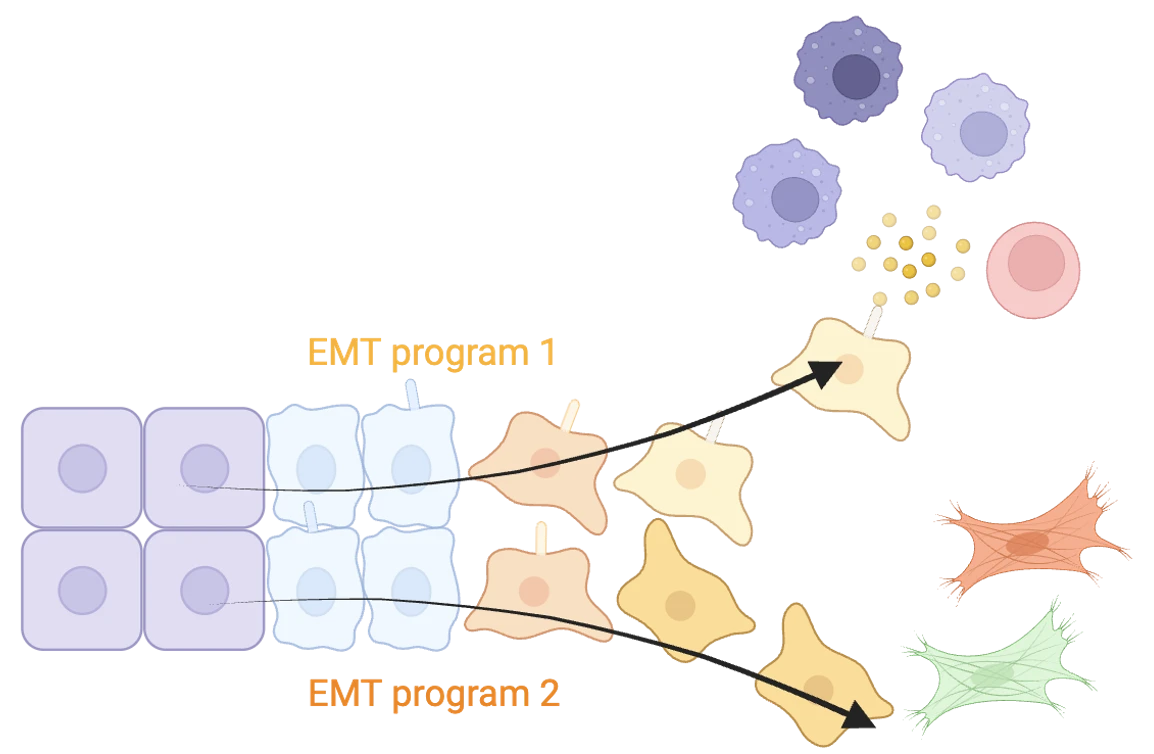
Research in the laboratory is now focused on mechanisms enabling cell state transitions to proceed along distinct branching pathways during the course of EMT rather than through a linear succession of cell states. We investigate the impact of branched evolution of cellular phenotypes on heterotypic cell-cell communications within cellular ecosystems both in normal and disease states. Our foundational research aims at revealing new vulnerabilities of diseases that can be targeted by innovative molecules of therapeutic interest that we develop for our research.
Tessier CE, Derrien J, Dupuy AMM, Pelé T, Moquet M, Roul J, Douillard E, El Harrif C, Pinson X, Le Gallo M, Godey F, Tas P, Viel R, Grasset E, Prigent C, Letouzé É, Suzanne P, Dallemagne P, Campone M, Weinberg RA, Lees JA, Juin PP, Guen VJ. EMT-ciliary signaling in quasi-mesenchymal-stem-like cells drives therapeutic resistance and is a druggable vulnerability in triple-negative breast cancer. EMBO Mol Med. 2025 Oct;17(10):2536-2561. doi:10.1038/s44321-025-00289-1. Epub 2025 Aug 26. PMID: 40859055; PMCID: PMC12514032.
Grasset EM, Deshpande A, Lee JW, Cho Y, Shin SM, Coyne EM, Hernandez A, Yuan X, Zhang Z, Cimino-Mathews A, Ewald AJ, Ho WJ. Mapping the breast tumor microenvironment: proximity analysis reveals spatial relationships between macrophage subtypes and metastasis-initiating cancer cells. Oncogene. 2024 Sep;43(39):2927-2937. doi: 10.1038/s41388-024-03127-9. Epub 2024 Aug 20. PMID: 39164522; PMCID: PMC12268993.
Grasset EM, Barillé-Nion S, Juin PP. Stress in the metastatic journey - the role of cell communication and clustering in breast cancer progression and treatment resistance. Dis Model Mech. 2024 Mar 1;17(3):dmm050542. doi: 10.1242/dmm.050542. Epub 2024 Mar 20. PMID: 38506114; PMCID: PMC10979546.
Tessier CE, Dupuy AMM, Pelé T, Juin PP, Lees JA, Guen VJ. EMT and primary ciliogenesis: For better or worse in sickness and in health. Genesis. 2024 Feb;62(1):e23568. doi: 10.1002/dvg.23568. Epub 2023 Nov 9. PMID: 37946671.
Dupuy AMM, Juin PP, Guen VJ. Using mammary organoids to study cilia. Methods Cell Biol. 2023;175:221-233. doi: 10.1016/bs.mcb.2022.09.010. Epub 2022 Nov 18. PMID: 36967142.
Grasset EM, Dunworth M, Sharma G, Loth M, Tandurella J, Cimino-Mathews A, Gentz M, Bracht S, Haynes M, Fertig EJ, Ewald AJ. Triple-negative breast cancer metastasis involves complex epithelial-mesenchymal transition dynamics and requires vimentin. Sci Transl Med. 2022 Aug 3;14(656):eabn7571. doi:10.1126/scitranslmed.abn7571. Epub 2022 Aug 3. PMID: 35921474; PMCID: PMC9801390.
Wilson MM, Callens C, Le Gallo M, Mironov S, Ding Q, Salamagnon A, Chavarria TE, Viel R, Peasah AD, Bhutkar A, Martin S, Godey F, Tas P, Kang HS, Juin PP, Jetten AM, Visvader JE, Weinberg RA, Attanasio M, Prigent C, Lees JA, Guen VJ. An EMT-primary cilium-GLIS2 signaling axis regulates mammogenesis and claudin-low breast tumorigenesis. Sci Adv. 2021 Oct 29;7(44):eabf6063. doi:10.1126/sciadv.abf6063. Epub 2021 Oct 27. PMID: 34705506; PMCID: PMC8550236.
Duclos M, Prigent C, Le Borgne R, Guen VJ. Three-Dimensional Imaging of Organoids to Study Primary Ciliogenesis During ex vivo Organogenesis. J Vis Exp. 2021 May 14;(171). doi: 10.3791/62365. PMID: 34057457.
Guen VJ, Prigent C. Targeting Primary Ciliogenesis with Small-Molecule Inhibitors. Cell Chem Biol. 2020 Oct 15;27(10):1224-1228. doi:10.1016/j.chembiol.2020.07.018. Epub 2020 Aug 13. PMID: 32795416.
Wilson MM, Weinberg RA, Lees JA, Guen VJ. Emerging Mechanisms by which EMT Programs Control Stemness. Trends Cancer. 2020 Sep;6(9):775-780. doi: 10.1016/j.trecan.2020.03.011. Epub 2020 Apr 17. PMID: 32312682.
Guen VJ, Edvardson S, Fraenkel ND, Fattal-Valevski A, Jalas C, Anteby I, Shaag A, Dor T, Gillis D, Kerem E, Lees JA, Colas P, Elpeleg O. A homozygous deleterious CDK10 mutation in a patient with agenesis of corpus callosum, retinopathy, and deafness. Am J Med Genet A. 2018 Jan;176(1):92-98. doi:10.1002/ajmg.a.38506. Epub 2017 Nov 12. PMID: 29130579; PMCID: PMC6190576.
Grasset EM, Bertero T, Bozec A, Friard J, Bourget I, Pisano S, Lecacheur M, Maiel M, Bailleux C, Emelyanov A, Ilie M, Hofman P, Meneguzzi G, Duranton C, Bulavin DV, Gaggioli C. Matrix Stiffening and EGFR Cooperate to Promote the Collective Invasion of Cancer Cells. Cancer Res. 2018 Sep 15;78(18):5229-5242. doi: 10.1158/0008-5472.CAN-18-0601. Epub 2018 Jul 19. PMID: 30026329.
Guen VJ, Chavarria TE, Kröger C, Ye X, Weinberg RA, Lees JA. EMT programs promote basal mammary stem cell and tumor-initiating cell stemness by inducing primary ciliogenesis and Hedgehog signaling. Proc Natl Acad Sci U S A. 2017 Dec 5;114(49):E10532-E10539. doi: 10.1073/pnas.1711534114. Epub 2017 Nov 20. PMID: 29158396; PMCID: PMC5724269.
Guen VJ, Gamble C, Perez DE, Bourassa S, Zappel H, Gärtner J, Lees JA, Colas P. STAR syndrome-associated CDK10/Cyclin M regulates actin network architecture and ciliogenesis. Cell Cycle. 2016;15(5):678-88. doi:10.1080/15384101.2016.1147632. PMID: 27104747; PMCID: PMC4846108.
Guen VJ, Gamble C, Flajolet M, Unger S, Thollet A, Ferandin Y, Superti-Furga A, Cohen PA, Meijer L, Colas P. CDK10/cyclin M is a protein kinase that controls ETS2 degradation and is deficient in STAR syndrome. Proc Natl Acad Sci U S A. 2013 Nov 26;110(48):19525-30. doi: 10.1073/pnas.1306814110. Epub 2013 Nov 11. PMID: 24218572; PMCID: PMC3845122.

Team leaders:
Vincent Guen CRCN Inserm, HDR
Eloïse Grasset CRCN CNRS
Team members:
Frédérique Nguyen, Associate Professor ONIRIS
Alice Serafin, Post-doctoral scientist
Alexandre Thomas, Post-doctoral scientist
Laurine Berland, PhD student
Chloé Grivaud, PhD student
Aurore Dupuy, PhD student
Jen Shapp, PhD student
Camille Tessier, PhD student
Lilian Chevreux, Master student
Quentin Batard, Engineer
Camille Hubert, Technician
Lab events

Institutions & Fundings

Contact us
We are open to applications from highly motivated individuals.
Vincent Guen, INSERM
vincent.guen@inserm.fr
Éloïse Grasset, CNRS
eloise.grasset@univ-nantes.fr