Team 07 : Stress Adaptation and Tumor Escape
Team leader: Dr Philippe P. Juin
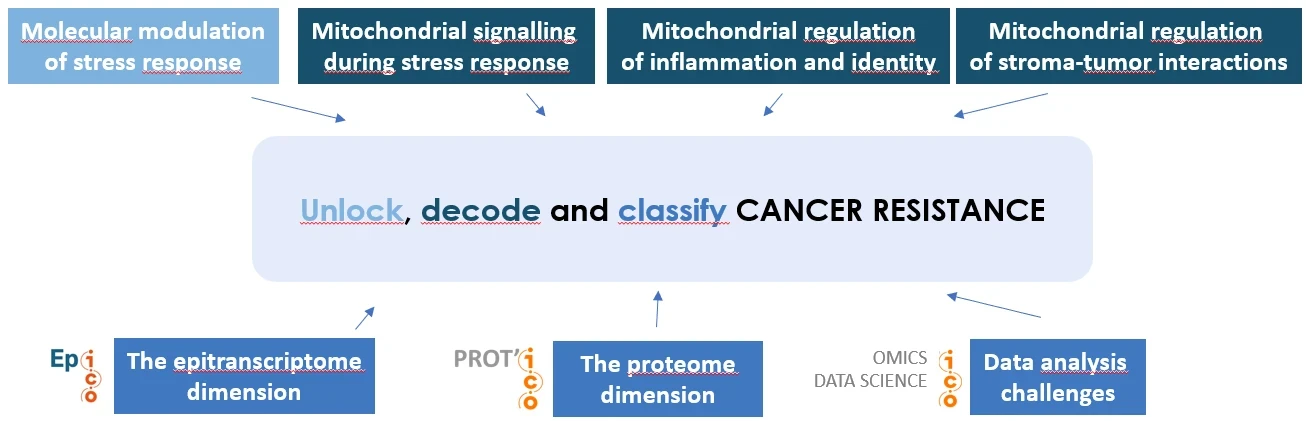
Stress detection and adaptation vary widely across tumors and cell populations, driving inter- and intra-patient heterogeneity. Because tumors are dynamically composed of diverse and plastic tumor states coevolving with the microenvironment, classifications based on dominant cell types at a given time is often insufficient: characterization of resistant tumor ecotypes relying on minor/transitory cell subpopulations or intercellular signals may escape standard comparative analysis. We thus develop analytical methodologies allowing to predict response and define groups of refractory tumors out of transcriptomic data, in strong link with ICO Bioinfomics Unit (P3, P5, P7). We also explore epitrancrisptomic programs, in particular those relying on miRNA and base modifications thereof, as they provide a critical layer coregulating numerous treatment responses, measurable in longitudinal liquid biopsies (P2, P10). This justifies our implication in research developments of the Ep’ICO platform complemented by pilot proteomic studies with the Prot’ICO platform.
Our connections with the clinics include implementation of cohorts, giving access to patient samples in order to functionnaly explore primary cells. A critical determinant of cell fate is mitochondrial integrity and we investigate it as part of an intercellular network influenced by the stroma. Cancer associated fibroblasts influence tumor cell mitochondrial integrity (P6) while their own mitochondrial integrity is key for their pathogenic phenotype (P4, P8). This underlines that, long regarded as a simple on/off switch controlling cell death in response to overwhelming stress, mitochondria function as a far more versatile signaling platform that regulates a broad spectrum of adaptive phenotypes. Relevantly, we have shown that the BCL-2 network of proteins, that guard mitochondrial integrity, respond to, and regulate, inflammatory phenotypes in primary breast cancers upon treatment (P1). We have unravelled key molecular features that regulate how these proteins function by engaging into protein complexes at subcellular membranes, opening new avenues for their pharmacological targeting (P9).
Our future project will fully leverage our expertise and build on these compelling results to uncover how stress responses are molecularly modulated, with a strong focus on mitochondrial signaling during stress. We will define the central role of mitochondria in shaping inflammation, tumor identity, and stroma–tumor interactions while further deploying innovative approaches to decode the molecular features of stress responses. We aim to better understand tumor escape, to help characterize treatment resistance and to circumscribe it by precision therapeutic strategies. (P1 to P10 and F1 to F10 refer to the major publications and fundings listed below)
|
|
You can read about our recent achievements in the text available at the top right.
Our Achievements 2020-2025
- Download the PDF
Latest news
 Laurent Maillet and co-authors are pleased to share their latest publication - well done! Click here to read more
Laurent Maillet and co-authors are pleased to share their latest publication - well done! Click here to read more A big thank you to the Ligue contre le Cancer for its long term support of the SATE team to support its research on the identification of MCL-1 dependent compensatory mechanisms contributing to cellular heterogeneity in aggressive breast cancers
A big thank you to the Ligue contre le Cancer for its long term support of the SATE team to support its research on the identification of MCL-1 dependent compensatory mechanisms contributing to cellular heterogeneity in aggressive breast cancers Philippe Juin, Winner of the Avenir 2022 Research Grant awarded by Association Ruban Rose for a project of collaboration between Team 7 and Team 12 on breast cancer Click here to watch the interview!
Philippe Juin, Winner of the Avenir 2022 Research Grant awarded by Association Ruban Rose for a project of collaboration between Team 7 and Team 12 on breast cancer Click here to watch the interview! 
P2. Cheray M, Etcheverry A, Jacques C, Pacaud R, Bougras-Cartron G, Aubry M, Denoual F, Peterlongo P, Nadaradjane A, Briand J, Akcha F, Heymann D, Vallette FM, Mosser J, Ory B, Cartron PF. Cytosine methylation of mature microRNAs inhibits their functions and is associated with poor prognosis in glioblastoma multiforme. Mol Cancer. 2020 Feb 25;19(1):36. doi: 10.1186/s12943-020-01155-z.PMID: 32098627
P3. Basseville A, Cordier C, Ben Azzouz F, Gouraud W, Lasla H, Panloup F, Campone M, Jézéquel P. Brain neural progenitors are new predictive biomarkers for breast cancer hormonotherapy. Cancer Research Communications. 2022 Jul. DOI: 10.1158/2767-9764.CRC-21-0090
P4. Bonneaud TL, Lefebvre CC, Nocquet L, Basseville A, Roul J, Weber H, Campone M, Juin PP, Souazé F. Targeting of MCL-1 in breast cancer-associated fibroblasts reverses their myofibroblastic phenotype and pro-invasive properties. Cell Death Dis. 2022 Sep 14;13(9):787. doi: 10.1038/s41419- 022-05214-9. PMID: 36104324; PMCID: PMC9474880.
P5. Jézéquel P, Lasla H, Gouraud W, Basseville A, Michel B, Frenel JS, Juin PP, Ben Azzouz F, Campone M. Mesenchymal-like immune-altered is the fourth robust triple-negative breast cancer molecular subtype Breast Cancer. 2024 Sep;31(5):825-840. doi: 10.1007/s12282-024-01597-z.
P6. Nocquet L, Roul J, Lefebvre CC, Duarte L, Campone M, Juin PP, Souazé F. Low BCL-xL expression in triple-negative breast cancer cells favors chemotherapy efficacy, and this effect is limited by cancer-associated fibroblasts. Sci Rep. 2024 Jun 19;14(1):14177. doi: 10.1038/s41598-024-64696-z. PMID: 38898061; PMCID: PMC11187150.
P7. Cordier C, Jézéquel P, Campone M, Panloup F, Basseville A. HABiC: an algorithm based on the exact computation of the Kantorovich-Rubinstein optimizer for binary classification in transcriptomics, Bioinformatics, 2025. DOI: 10.1093/bioinformatics/btaf310
P8. Lefebvre CC, Giowachini P, Derrien J, Naour M, Corre I, Thirouard L, Douillard E, Chiron D, GuillonneauF, Treps L, Campone M, Juin PP, Souazé F. MCL-1 as a molecular switch between myofibroblastic and pro-angiogenic features of breast cancer-associated fibroblasts. Cell Death Dis. 2025 Aug 9;16(1):603. doi: 10.1038/s41419-025-07920-6. PMID: 40783386; PMCID: PMC12335522.
P9. Maillet L, Fétiveau A, Lalier L, Martin N, Barillé-Nion S, Guette C, Gautier F, Téletchéa S, Juin PP. Allosteric regulation of BH3-in-groove interactions by tail anchors of BCL-xL complexes limits BH3 mimetic antagonism. Nature Commun. 2025 doi.org/10.1038/s41467-025-65509-1
P10.Besson JM, Etcheverry A, Nadaradjane A, Bougras-Cartron G, Aubry M, Frenel JS, Chevanieu A, Lopez M, Mosser J, Cartron PF. Selective disruption of DNMT1/ELK1 interactions induces DGKI re-expression and promotes temozolomide sensitivity of MGMTmethylated/DGKImethylatedglioblastoma.Clin Epigenetics. 2025 Aug 30;17(1):146. doi: 10.1186/s13148-025-01943-8. PMID: 40886022
2026
2025
- Maillet L, Fétiveau A, Lalier L, Martin N, Barillé-Nion S, Guette C, Gautier F, Téletchéa S, Juin PP. Allosteric regulation of BH3-in-groove interactions by tail anchors of BCL-xL complexes limits BH3 mimetic antagonism. Nat Commun. 2025 Nov 22. doi: 10.1038/s41467-025-65509-1. Epub ahead of print. PMID: 41274891.
- Besson JM, Etcheverry A, Nadaradjane A, Bougras-Cartron G, Aubry M, Frenel JS, Chevanieu A, Lopez M, Mosser J, Cartron PF. Selective disruption of DNMT1/ELK1 interactions induces DGKI re-expression and promotes temozolomide sensitivity of MGMTmethylated/DGKImethylated glioblastoma. Clin Epigenetics. 2025 Aug 30;17(1):146. doi: 10.1186/s13148-025-01943-8. PMID: 40886022; PMCID: PMC12398979.
- Tessier CE, Derrien J, Dupuy AMM, Pelé T, Moquet M, Roul J, Douillard E, El Harrif C, Pinson X, Le Gallo M, Godey F, Tas P, Viel R, Grasset E, Prigent C, Letouzé É, Suzanne P, Dallemagne P, Campone M, Weinberg RA, Lees JA, Juin PP, Guen VJ. EMT-ciliary signaling in quasi-mesenchymal-stem-like cells drives therapeutic resistance and is a druggable vulnerability in triple-negative breast cancer. EMBO Mol Med. 2025 Oct;17(10):2536-2561. doi: 10.1038/s44321-025-00289-1. Epub 2025 Aug 26. PMID: 40859055; PMCID: PMC12514032
- Lefebvre CC, Giowachini P, Derrien J, Naour M, Corre I, Thirouard L, Douillard E, Chiron D, Guillonneau F, Treps L, Campone M, Juin PP, Souazé F. MCL-1 as a molecular switch between myofibroblastic and pro-angiogenic features of breast cancer-associated fibroblasts.Cell Death Dis. 2025 Aug 9;16(1):603. doi: 10.1038/s41419-025-07920-6.PMID: 40783386
- Dilasser F, Rose L, Quemener A, Ferrandez Y, Hassoun D, Rousselle M, Bergereau H, Lambot SM, Anselmino LE, Trouillet C, Andre G, Maillasson M, Croyal M, Riviere M, Dubreuil D, Collet S, Souaze F, Campone M, Patsouris A, Mortier E, Marquez MM, Juin P, Lebreton J, Tessier A, Cherfils J, Loirand G, Sauzeau V. A Rac-specific competitive inhibitor of guanine nucleotide binding reduces metastasis in triple-negative breast cancer. Cell Rep Med. 2025 Jul 15;6(7):102233. doi: 10.1016/j.xcrm.2025.102233. Epub 2025 Jul 8. PMID: 40633540; PMCID: PMC12281424..
- Gaber M, Quentel A, Holmes J, Lepetit C, Triki H, Wilson A, Payne V, Tenvooren I, Dehours C, Peoples A, Duet ML, Katz AJ, Pécot T, Bougras-Cartron G, Cartron PF, Cook KL, Vidi PA. Obesity increases DNA damage in the breast epithelium. Breast Cancer Res. 2025 Jan 21;27(1):11. doi: 10.1186/s13058-025-01961-7. PMID: 39838489; PMCID: PMC11753040.
2024
- Guen VJ, André-Grégoire G, Beauvillain C, Boury F, Chauvet M, Dupuy AMM, Fonteneau JF, Gagne K, Gavard J, Gomez-Bougie P, Grasset E, Jardine J, Lamoureux F, Laurent-Blond M, Letouzé É, Macé Y, Maurice S, Pecqueur C, Pouliquen D, Rbah-Vidal L, Pellat-Deceunynck C, Juin PP. CRCI2NA inaugural symposium: A meeting on tumor and immune ecosystems. Biol Cell. 2024 Nov 9:e202400111. doi: 10.1111/boc.202400111. Epub ahead of print. PMID: 39520372.
- Nocquet L, Roul J, Lefebvre CC, Duarte L, Campone M, Juin PP, Souazé F. Low BCL-xL expression in triple-negative breast cancer cells favors chemotherapy efficacy, and this effect is limited by cancer-associated fibroblasts. Sci Rep. 2024 Jun 19;14(1):14177. doi: 10.1038/s41598-024-64696-z. PMID: 38898061; PMCID: PMC11187150.
- Jézéquel P, Lasla H, Gouraud W, Basseville A, Michel B, Frenel JS, Juin PP, Ben Azzouz F, Campone M. Mesenchymal-like immune-altered is the fourth robust triple-negative breast cancer molecular subtype. Breast Cancer. 2024 May 22. doi: 10.1007/s12282-024-01597-z. Epub ahead of print. PMID: 38777987.
- Grasset EM, Barillé-Nion S, Juin PP. Stress in the metastatic journey - the role of cell communication and clustering in breast cancer progression and treatment resistance. Dis Model Mech. 2024 Mar 1;17(3):dmm050542. doi: 10.1242/dmm.050542. Epub 2024 Mar 20. PMID: 38506114; PMCID: PMC10979546
- Tessier CE, Dupuy AMM, Pelé T, Juin PP, Lees JA, Guen VJ. EMT and primary ciliogenesis: For better or worse in sickness and in health. Genesis. 2024 Feb;62(1):e23568. doi: 10.1002/dvg.23568. Epub 2023 Nov 9. PMID: 37946671
2023
- Dupuy AMM, Juin PP, Guen VJ. Using mammary organoids to study cilia. Methods Cell Biol. 2023;175:221-233. doi: 10.1016/bs.mcb.2022.09.010. PMID: 36967142.
- Lavy M, Gauttier V, Dumont A, Chocteau F, Deshayes S, Fresquet J, Dehame V, Girault I, Trilleaud C, Neyton S, Mary C, Juin P, Poirier N, Barillé-Nion S, Blanquart C. ChemR23 activation reprograms macrophages toward a less inflammatory phenotype and dampens carcinoma progression. Front Immunol. 2023 Jul 19;14:1196731. doi: 10.3389/fimmu.2023.1196731. PMID: 37539056;
- Bougras-Cartron G, Nadaradjane A, Joalland MP, Lalier-Bretaudeau L, Raimbourg J, Cartron PF. Adenosine Methylation Level of miR-125a-5p Promotes Anti-PD-1 Therapy Escape through the Regulation of IGSF11/VSIG3 Expression. Cancers (Basel). 2023 Jun 14;15(12):3188. doi: 10.3390/cancers15123188. PMID: 37370798;
- Bikfalvi A, da Costa CA, Avril T, Barnier JV, Bauchet L, Brisson L, Cartron PF, Castel H, Chevet E, Chneiweiss H, Clavreul A, Constantin B, Coronas V, Daubon T, Dontenwill M, Ducray F, Enz-Werle N, Figarella-Branger D, Fournier I, Frenel JS, Gabut M, Galli T, Gavard J, Huberfeld G, Hugnot JP, Idbaih A, Junier MP, Mathivet T, Menei P, Meyronet D, Mirjolet C, Morin F, Mosser J, Moyal EC, Rousseau V, Salzet M, Sanson M, Seano G, Tabouret E, Tchoghandjian A, Turchi L, Vallette FM, Vats S, Verreault M, Virolle T. Challenges in glioblastoma research: focus on the tumor microenvironment. Trends Cancer. 2023 Jan;9(1):9-27. doi: 10.1016/j.trecan.2022.09.005.
2022
- Courant F, Bougras-Cartron G, Abadie C, Frenel JS, Cartron PF. Modulation of DNA Methylation/Demethylation Reactions Induced by Nutraceuticals and Pollutants of Exposome Can Promote a C > T Mutation in the Breast Cancer Predisposing Gene PALB2. Epigenomes. 2022 Sep 30;6(4):32. doi: 10.3390/epigenomes6040032. PMID: 36278678
- Wilson MM, Callens C, Le Gallo M, Mironov S, Ding Q, Salamagnon A, Chavarria TE, Viel R, Peasah AD, Bhutkar A, Martin S, Godey F, Tas P, Kang HS, Juin PP, Jetten AM, Visvader JE, Weinberg RA, Attanasio M, Prigent C, Lees JA, Guen VJ. An EMT-primary cilium-GLIS2 signaling axis regulates mammogenesis and claudin-low breast tumorigenesis. Sci Adv. 2021 Oct 29;7(44):eabf6063.
- Bonneaud TL, Lefebvre CC, Nocquet L, Basseville A, Roul J, Weber H, Campone M, Juin PP, Souazé F. Targeting of MCL-1 in breast cancer-associated fibroblasts reverses their myofibroblastic phenotype and pro-invasive properties. Cell Death Dis. 2022 Sep 14;13(9):787. doi: 10.1038/s41419-022-05214-9.
- Lalier L, Vallette F, Manon S. Bcl-2 Family Members and the Mitochondrial Import Machineries: The Roads to Death. Biomolecules. 2022 Jan 19;12(2):162.
- Garnier D, Ratcliffe E, Briand J, Cartron PF, Oliver L, Vallette FM. The Activation of Mesenchymal Stem Cells by Glioblastoma Microvesicles Alters Their Exosomal Secretion of miR-100-5p, miR-9-5p and let-7d-5p. Biomedicines. 2022 Jan 6;10(1):112.
2021
- Wilson MM, Callens C, Le Gallo M, Mironov S, Ding Q, Salamagnon A, Chavarria TE, Viel R, Peasah AD, Bhutkar A, Martin S, Godey F, Tas P, Kang HS, Juin PP, Jetten AM, Visvader JE, Weinberg RA, Attanasio M, Prigent C, Lees JA, Guen VJ. An EMT-primary cilium-GLIS2 signaling axis regulates mammogenesis and claudin-low breast tumorigenesis. Sci Adv. 2021 Oct 29;7(44):eabf6063.
- Hazan R, Mori M, Danielian PS, Guen VJ, Rubin SM, Cardoso WV, Lees JA. E2F4's cytoplasmic role in multiciliogenesis is mediated via an N-terminal domain that binds two components of the centriole replication machinery, Deup1 and SAS6. Mol Biol Cell. 2021 Oct 1;32(20):ar1.
- Lavy M, Gauttier V, Poirier N, Barillé-Nion S, Blanquart C. Specialized Pro-Resolving Mediators Mitigate Cancer-Related Inflammation: Role of Tumor-Associated Macrophages and Therapeutic Opportunities. Front. Immunol., 30 June 2021 | https://doi.org/10.3389/fimmu.2021.702785.
- Tea I, De Luca A, Schiphorst AM, Grand M, Barillé-Nion S, Mirallié E, Drui D, Krempf M, Hankard R, Tcherkez G. Stable Isotope Abundance and Fractionation in Human Diseases. Metabolites. 2021 Jun 9;11(6):370.
- Duclos M, Prigent C, Le Borgne R, Guen VJ. Three-Dimensional Imaging of Organoids to Study Primary Ciliogenesis During ex vivo Organogenesis. J Vis Exp. 2021 May 14;(171).
- Jézéquel P, Gouraud W, Ben Azzouz F, Guérin-Charbonnel C, Juin PP, Lasla H, Campone M. bc-GenExMiner 4.5: new mining module computes breast cancer differential gene expression analyses. Database (Oxford). 2021 Feb 18; 2021:baab007. doi: 10.1093/database/baab007.
- Ben Azzouz F, Michel B, Lasla H, Gouraud W, François AF, Girka F, Lecointre T, Guérin-Charbonnel C, Juin PP, Campone M, Jézéquel P. Development of an absolute assignment predictor for triple-negative breast cancer subtyping using machine learning approaches. Comput Biol Med. 2021 Feb;129:104171. doi: 10.1016/j.compbiomed.2020.104171.
- Lalier L, Mignard V, Joalland MP, Lanoé D, Cartron PF, Manon S, Vallette FM. TOM20-mediated transfer of Bcl2 from ER to MAM and mitochondria upon induction of apoptosis. Cell Death Dis. 2021 Feb 15;12(2):182.
- Olivier C, Oliver L, Lalier L, Vallette FM. Drug Resistance in Glioblastoma: The Two Faces of Oxidative Stress. Front Mol Biosci. 2021 Jan 27;7:620677.
2020
- Guyon N, Garnier D, Briand J, Nadaradjane A, Bougras-Cartron G, Raimbourg J, Campone M, Heymann D, Vallette FM, Frenel JS, Cartron PF. Anti-PD1 therapy induces lymphocyte-derived exosomal miRNA-4315 release inhibiting Bim-mediated apoptosis of tumor cells. Cell Death Dis. 2020 Dec 11;11(12):1048. doi: 10.1038/s41419-020-03224-z.
- Nocquet L, Juin PP, Souazé F. Mitochondria at Center of Exchanges between Cancer Cells and Cancer-Associated Fibroblasts during Tumor Progression. Cancers (Basel). 2020 Oct 17;12(10):3017. doi: 10.3390/cancers12103017.
- Guen VJ, Prigent C. Targeting Primary Ciliogenesis with Small-Molecule Inhibitors. Cell Chem Biol. 2020 Oct 15;27(10):1224-1228.
- Peixoto P, Cartron PF, Serandour AA, Hervouet E. From 1957 to Nowadays: A Brief History of Epigenetics. Int J Mol Sci. 2020 Oct 14;21(20):7571. doi: 10.3390/ijms21207571. PMID: 33066397;
- Wilson MM, Weinberg RA, Lees JA, Guen VJ. Emerging Mechanisms by which EMT Programs Control Stemness. Trends Cancer. 2020 Sep;6(9):775-780.
- Briand J, Sérandour AA, Nadaradjane A, Bougras-Cartron G, Heymann D, Ory B, Vallette FM, Cartron PF. N6-Adenosine Methylation of miRNA-200b-3p Influences Its Functionality and Is a Theranostic Tool. Mol Ther Nucleic Acids. 2020 Aug 14;22:72-83. doi: 10.1016/j.omtn.2020.08.010.
- Oliver L, Lalier L, Salaud C, Heymann D, Cartron PF, Vallette FM. Drug resistance in glioblastoma: are persisters the key to therapy? Cancer Drug Resist. 2020 Aug 7;3(3):287-301. doi: 10.20517/cdr.2020.29.
- Mignard V, Dubois N, Lanoé D, Joalland MP, Oliver L, Pecqueur C, Heymann D, Paris F, Vallette FM, Lalier L. Sphingolipid distribution at mitochondria-associated membranes (MAMs) upon induction of apoptosis. J Lipid Res. 2020 Jul;61(7):1025-1037.
- Barillé-Nion S, Lohard S, Juin PP: Targeting of BCL-2 Family Members during Anticancer Treatment: A Necessary Compromise between Individual Cell and Ecosystemic Responses? Biomolecules 2020 Jul 25;10(8):E1109. doi: 10.3390/biom10081109.
- Lohard S, Juin PP, Barillé-Nion S. Mitotic stress-induced secretome primes cancer cells to apoptosis and maximizes paclitaxel response in breast tumors when combined with BCL-xL-targeting BH3 mimetics. Mol Cell Oncol. 2020 Mar 19;7(3):1735912. doi: 10.1080/23723556.2020.1735912.
- Briand J, Garnier D, Nadaradjane A, Clément-Colmou K, Potiron V, Supiot S, Bougras-Cartron G, Frenel JS, Heymann D, Vallette FM, Cartron PF. Radiotherapy-induced overexpression of exosomal miRNA-378a-3p in cancer cells limits natural killer cells cytotoxicity. Epigenomics. 2020 Mar;12(5):397-408. doi: 10.2217/epi-2019-0193.
- Cheray M, Etcheverry A, Jacques C, Pacaud R, Bougras-Cartron G, Aubry M, Denoual F, Peterlongo P, Nadaradjane A, Briand J, Akcha F, Heymann D, Vallette, FM, Mosser J, Ory B, Cartron PF. Cytosine methylation of mature microRNAs inhibits their functions and is associated with poor prognosis in glioblastoma multiforme. Mol Cancer. 2020 Feb 25;19(1):36. doi: 10.1186/s12943-020-01155-z.
- Lohard S, Bourgeois N, Maillet L, Gautier F, Fétiveau A, Lasla H, Nguyen F, Vuillier C, Dumont A, Moreau-Aubry A, Frapin M, David L, Loussouarn D, Kerdraon O, Campone M, Jézéquel P, Juin PP, Barillé-Nion S. STING-dependent paracriny shapes apoptotic priming of breast tumors in response to anti-mitotic treatment. Nature Communications. 2020 11:259. doi.org/10.1038/s41467-019-13689-y
- Duforestel M, Briand J, Bougras-Cartron G, Heymann D, Frenel JS, Vallette FM, Cartron PF. Cell-free circulating epimarks in cancer monitoring. Epigenomics. 2020 Jan;12(2):145-155. doi: 10.2217/epi-2019-0170.
- Cartron PF, Cheray M, Bretaudeau L. Epigenetic protein complexes: the adequate candidates for the use of a new generation of epidrugs in personalized and precision medicine in cancer. Epigenomics. 2020 Jan;12(2):171-177. doi: 10.2217/epi-2019-0169.
- Rabé M, Dumont S, Álvarez-Arenas A, Janati H, Belmonte-Beitia J, Calvo GF, Thibault-Carpentier C, Séry Q, Chauvin C, Joalland N, Briand F, Blandin S, Scotet E, Pecqueur C, Clairambault J, Oliver L, Perez-Garcia V, Nadaradjane A, Cartron PF, Gratas C, Vallette FM. Identification of a transient state during the acquisition of temozolomide resistance in glioblastoma. Cell Death Dis. 2020 Jan 6;11(1):19.
2019
- Avril P, Vidal L, Barille-Nion S, Le Nail LR, Redini F, Layrolle P, Pinault M, Chevalier S, Perrot P, Trichet V. Epinephrine Infiltration of Adipose Tissue Impacts MCF7 Breast Cancer Cells and Total Lipid Content. Int J Mol Sci. 2019 Nov 11;20(22). pii: E5626.
- Briand J, Nadaradjane A, Bougras-Cartron G, Olivier C, Vallette FM, Cartron PF. Diuron exposure and Akt overexpression promote glioma formation through DNA hypomethylation. Clin Epigenetics. 2019 Nov 14;11(1):159. doi: 10.1186/s13148-019-0759-1.
- Duforestel M, Nadaradjane A, Bougras-Cartron G, Briand J, Olivier C, Frenel JS, Vallette FM, Lelièvre SA, Cartron PF. Glyphosate Primes Mammary Cells for Tumorigenesis by Reprogramming the Epigenome in a TET3-Dependent Manner. Front Genet. 2019 Sep 27;10:885. doi: 10.3389/fgene.2019.00885.
- Vallette FM, Olivier C, Lézot F, Oliver L, Cochonneau D, Lalier L, Cartron PF, Heymann D. Dormant, quiescent, tolerant and persister cells: Four synonyms for the same target in cancer. Biochem Pharmacol. 2019 Apr;162:169-176. doi: 10.1016/j.bcp.2018.11.004.
- Louault K, Bonneaud TL, Séveno C, Gomez-Bougie P, Nguyen F, Gautier F, Bourgeois N, Loussouarn D, Kerdraon O, Barillé-Nion S, Jézéquel P, Campone M, Amiot M, Juin PP, Souazé F. Interactions between cancer-associated fibroblasts and tumor cells promote MCL-1 dependency in estrogen receptor-positive breast cancers. Oncogene. 2019 Apr;38(17):3261-3273. doi: 10.1038/s41388-018-0635-z.
- Brown HK, Tellez-Gabriel M, Cartron PF, Vallette FM, Heymann MF, Heymann D. Characterization of circulating tumor cells as a reflection of the tumor heterogeneity: myth or reality? Drug Discov Today. 2019 Mar;24(3):763-772. doi: 10.1016/j.drudis.2018.11.017.
- Blanquart C, Linot C, Cartron PF, Tomaselli D, Mai A, Bertrand P. Epigenetic Metalloenzymes. Curr Med Chem. 2019;26(15):2748-2785.
- Ramana Murthy AV, Narendar V, Kumar NS, Aparna P, Durga Bhavani AK, Gautier F, Barillé-Nion S, Juin P, Mosset P, Grée R, Levoin N. Targeting PUMA/Bcl-xL interaction by new specific compounds to unleash apoptotic process in cancer cells. Eur J Med Chem. 2019 Jan 15;162:334-347.
Sophie Barillé-Nion (CRHC INSERM, Nantes 1)
Agnès Basseville (Res. ICO, Angers, 0,5)
Pierre-François Cartron (CRCN INSERM, St Herblain, 1)
Jennifer Derrien (Res., ICO, St Herblain and Angers, 0,4)
Fabien Gautier (Res. ICO, St Herblain 0,8)
François Guillonneau (Res. ICO, Angers 0.5)
Lisenn Lalier (Res. ICO , St Herblain, 0.8)
Laurent Maillet (CRHC INSERM, Nantes, 1)
Aurélien Sérandour (MC ECN, St Herblain, 0.5)
Frédérique Souazé (CRHC Inserm, Nantes, 1)
Mario Campone (PU/PH, ICO, St Herblain and Angers, 0.3)
Gwenola Cartron (Project manager ICO, St Herblain 0.8)
Mathilde Dupé (PH ICO St Herblain 0.2)
Jean-Sébastien Frenel (Oncologist, ICO St Herblain 0.3)
Pascal Jezequel (PH, ICO, St Herblain 0.2)
Judith Raimbourg (PH, ICO, St Herblain 0.3)
Olivier Kerdraon (PH, ICO, St Herblain 0.2)
Thomas Cabioch (tech ICO, St Herblain, molecular and cellular biology, biochemistry 1)
Aurélie Fétiveau (tech Nantes Université, Nantes,molecular biology including constructs, cellular biology including plasmid 3D culture maintenance, biochemistry, genome editing, BRET assays, time lapse microscopy, Prevention Assistant 1)
Arulraj Nadaradjane (tech Nantes université, St Herblain, 2D culture, molecular biology related to single cell and long-read sequencing, Prevention Assistant, 1)
Julie Roul (tech ICO, Nantes, patient sample handling, primary 3D cultures, biochemistry, cytometry and single cell emulsion, 1)
Non-permanent members
Jean-Maxime Besson (Post-doc, St Herblain, 2024-2026, 1)Claire Lalevée (PhD student, St Herblain, 2024-2027, 1)
Sarah Kamami (PhD student, Angers, 2025-2028, 1)
Doran Espi-Bastien (tech St Herblain, cell biology and biochemistry in ovo assays, 2023-2026,1)
Eva Denion (Engineer Angers, proteomic manipulations 2025-2026, 1)
Laurine Duarte (IE Nantes, 2D and 3D cultures, immune assays, cytometry, time lapse microscopy, metabolism 2026-2027, 1)
Vanessa Josso (IE Nantes, 2D and 3D cultures, cytometry, biochemical assays including BioID, animal experimentations 2024-2026, 1)
Néna Martin (Engineer, Nantes, molecular and cellular biology, biochemistry, genome editing, time lapse microscopy 2025 -2027, 1)
Total FTEs: 23.3

Training, highlight number of HDR, PhD students trained (2020-2025), teaching activities
T1. Eight team members have the “Habilitation à Diriger les Recherches” (HDR)
T2. Post-doc mentoring (4) include H Triki (2023-2024) and JM Besson (since 2025) for PF Cartron and V Guen (2021-2022) and E Grasset (2023-2024) for PP Juin
T3. PhD mentoring (9 sustained) include M Lavy (2021, with C Blanquart Team 1) A Dumont (2023), F Chocteau (2025) for S Barillé; C Cordier (2025) for A Basseville; T Bonneaud (2021), L Nocquet (2023), C Lefebvre (2025) for F Souazé; M Duforestel (2020), F Courant (2022) for PF Cartron.
T4. Teaching activities are as follows: S Barillé: UE Recherche DFGSM2 (Médecine-Nantes Univ/2H); A Basseville: Master2 parcours ICAT, UE innovations thérapeutiques, UA (2h/year); PF Cartron: Master2 SANH and GP-OHNU (Nantes université, 4h/year); PP Juin Master 2 GP-OHNU (2h/year); L Maillet: Master2 Biochimie University of Bordeaux (4h/year)
CURRENT
Sarah Kamami: Caractérisation phénotypique du cancer du sein par profilage protéomique stochastiqueClaire Lalevée: Identification d’opportunités thérapeutiques sur la base du décryptage des méthylomes et du transcriptome des tumeurs
Laurine Berland: EMBRAVE-TNBC: Exploring Mechanisms of Breast Cancer Metastasis through Epithelial-Mesenchymal Transition and Vimentin Expression in Triple-Negative Breast Cancer
Camille Tessier: Development of small-molecule inhibitors of primary ciliogenesis for research on primary cilia
Aurore Dupuy: Investigating the role of the primary cilium in mammary gland remodeling during the reproduction cycle
PAST
Chloé Lefebvre : Role of MCL-1 in myofibroblast phenotype and chemoresistance of fibroblasts associated with breast cancers. Defense on September 16, 2025
Florian Chocteau: Metastatic process in breast cancer after chemotherapy treatment: Immune microenvironment and tumor clonal evolution in the metastatic niche. Defense on February 25, 2025
Alison Dumont: Heterogeneity in the response of breast tumors to antimitotics: What impacts on treatment resistance and immune response? Defense on December 11, 2023
Lisa Nocquet: Implication of BCL-xL and MCL-1 in the metabolic control of cell death by cancer-associated fibroblasts in triple negative breast cancer. Defense on November 30, 2023
Florestan Courant: Cross-species differences or similarities in epigenetic mechanisms induced by pesticides or chemotherapeutic agents. Defense on October 17, 2022
Nina Belaid: Modulation of KRAS activity by BCL-xL: mechanistic aspects and therapeutic implications. Defense on May 18, 2022
Thomas Bonneaud: Consequences of pharmacological targeting of MCL-1 on the myofibroblastic phenotype of breast cancer-associated fibroblasts. Defense on December 3, 2021
Margot Lavy: Regulating myeloid cell activity in tumors through Resolvlne receptors : chemR23. Defense on December 13, 2021
Manon Duforestel: Study of epigenetic mechanisms and dynamics in glioblastoma multiforme during the acquisition of temozolomide resistance. Defense on September 17, 2021
Steven Lohard: STING-dependent paracriny shapes apoptotic priming of breast tumors in response to antimitotic treatment. Defense on May 3, 2019
Josephine Briand: Glioblastoma and epigenetics: from prevention to development of new treatments. Defense on December 12, 2019
Kevin Louault: Role of Cancer-associated fibroblast in apoptosis resistance in breast cancers. Defense on December 19, 2018
Céline Vuillier: Mitochondrial apoptosis regulated by E2F1. Involvement of the E2F1 mitochondrial targeting and interactions with the Bcl-2 Family anti-apoptotic members. Defense on December 19, 2016
Jessie Pécot: BCLxL dependence of cancer cells. Targeting of the dynamic BCLxL/PUMA/BAX network. Oncogenic pathways regulated by the non canonical RAS/BCLxL interaction (October 2015)
Eloise Véquaud: Study of mammary cancer cells response to silencing of surviving by RNA interference or pharmacological targeting: highlighting oh homologous recombination regulation by survivin. (December 2014)
From 2019 Thomas Courouble 2025/26 : Master 2 Biologie-Santé, Université de Reims Champagne-Ardenne, Modélisation moléculaire de l'interaction MCL-1 / NOXA en contexte membranaire
Farida Guira 2025/26 : Master 2 OHNU, Quel rôle de NFkB dans la signalisation de la voie cGAS-STING induite par le taxol dans les TNBC?
Justine Philippe 2025/26 : Master 2 Biologie cellulaire et moléculaire, Université de Rennes, France. Influence de la matrice extra-cellulaire généré par les CAFs sur le phénotype et la chimiorésistance des cellules cancéreuses mammaires
Tom Moraitis 2024/25 : Master 2 Recherche mention « Biologie Santé » parcours Oncologie, immunologie, génétique (OIG). Décryptage d’anomalies épigénétiques/épitranscriptomiques/transcriptomiques associées à la résistance de traitements anti-tumoraux en opportunités thérapeutiques. UFR Sciences et Techniques de l’Université de Rouen Normandie
Marie Gonçalves 2024/25 : Master 2 TOXICOLOGIE ET ECOTOXICOLOGIE PARCOURS TOXICOLOGIE HUMAINE EVALUATION DES RISQUES ET VIGILANCES, Université Paris Cité. Caractérisation de la réponse au stress induit par un ciblage mitochondrial sous influence des fibroblastes associés au cancer pour améliorer la réponse au traitement des cellules cancéreuses du sein triple négatif
Mounia Elhannani 2024/25 : Master 2 Graduate Program Oncologie, Hématologie et médecine NUcléaire (GP-OHNU), Nantes Université, France. Rôle des CAFs dans l’élaboration de matrices extra-cellulaires pro-tumorales et la chimiorésistance des cancers du sein
Quentin Le Borgne 2024/25 : Master 2 I3, Nantes Université, France
Abed-el-Razzak Saleh 2024/25 : Master 2 Graduate Program Oncologie, Hématologie et médecine NUcléaire (GP-OHNU), Nantes Université, France
Morgan Zenatri 2023/24 : Master 2 Biologie-Santé, Nantes Université, France.
Angèle Palierne 2023/24 : Master 2 Biologie-Santé, Nantes Université, France. Etude des mécanismes cellulaires sous-jacents à la résistance aux traitements des cancers du sein induite par les fibroblastes du microenvironnement tumoral
Flavie Bernigole 2023/24 : Master 2 Génétique, Génomique et Biologie des Système, Nantes Université, France. Étude de l'hétérogénéité cellulaire dans la glande mammaire au cours du cycle de reproduction
Claire Lalevée 2023/24 : Master 2 Génétique, Génomique et Biologie des Système, Nantes Université, France. Analyse des profils cellulaires résistants par transcriptomique spatiale dans les cancers du sein réfractaires aux traitements
Audrey Roussel 2022/2023: M2 Biologie Moléculaire et Cellulaire Université de Rennes - Définir l'interactome de MCL-1 par BioID APEX
Mathilde Dupé 2022/23: Master 2 Biologie -Santé (Interactions Cellulaires et Applications Thérapeutiques ) University of Angers, France.Angers. Highlighting therapeutic opportunities induced by studying miRome reprogramming during the development of anastrozole resistance in endometrial cancer.
Aristide Massé 2022/23: Master2 BBRT, University of Nantes, France. Investigating the kinetics of MAM and the impact in the mitochondrial adaptation and resistance acquisition of cancer cells
Camille Tessier 2022/23 : Master 2 Biologie-Santé, Nantes Université, France. Investigating the role of epithelial plasticity and of primary cilia in breast cancer therapeutic resistance
Nicolas Balloud 2022/23: M2 BBRT (Biologie Biotechnologie Recherche Thérapeutique) University of Nantes, France. Deciphering the epigenetic/epitranscriptomic reprogramming associated with acquisition of tolerance/resistance to targeted antitumor therapies
Aurore Dupuy 2021/22: Master 2 "Biologie Moléculaire et Cellulaire", University of Rennes 1, France. Investigating the role of the primary cilium in mammary gland remodeling during the reproduction cycle.
Maxime Richomme 2021/22: Master 2 BBRT, Nantes University
Klara Edern 2021/22: Equivalent Master 2 Bioinformatics at Ecole Centrale de Nantes "High throughput data analysis to predict tumor cell fate and/or the existence of therapeutic vulnerabilities"
Morgane Allory 2021/22: Master 2 Biologie Santé parcours "Interactions cellulaires et Application thérapeutiques", Angers University. Identification of epigentic and/or epitranscriptomic biomarkers predictive of vulnerabilities to drugs/targeted therapy molecules.
Chloé Lefebvre 2021/22: Master 2 "Biologie, Biotechnologie et Recherche Thérapeutique", Nantes University. Role of MCL-1 in myofibroblast phenotype and chemoresistance of fibroblasts associated with breast cancers.
Hugo Weber 2020/21: Master 2 « Oncologie Moléculaire et Biothérapies », Limoges University. Effect of irradiation on the sensitivity of CAFs and breast adenocarcinoma cancer cells to BH3 mimetics: Evaluation of their therapeutic potential as senolytic agents.
Yuna Landais 2020/2021: Equivalent Master 2 Bioinformatics at Ecole Centrale de Nantes. "Study of mitochondrial genome methylation by direct nanopore sequencing"
Elen Goujon 2020/2021: Equivalent Master 2 Bioinformatics at Ecole Centrale de Nantes. "Development of biomedical data analysis pipelines in oncology"
Piera Grisolina 2019/2020: ERASMUS. Master 2 "Biologie Moléculaire et Cellulaire" University of Rennes 1/Tieste University, Italy. Investigating the molecular mechanisms by which EMT programs induce primary ciliogenesis.
Alison Dumont 2019/2020: Master 2 "Biologie, Biotechnologie et Recherche Thérapeutique", Nantes University: Induction of NOXA pro-apoptotic protein expression in response to Paclitaxel in breast tumors: Regulome analysis of its gene transcription.
Louis Paré 2019/2020: Equivalent Master 2 Bioinformatics at Ecole Centrale de Nantes. "Cinetic study of the resistance of anti-tumor treatments by epigenetic genome analysis"
Master 2
No specific announcement at the moment.PhD Thesis
No specific announcement at the moment.
Post docs
No specific announcement at the moment.Others job opportunities
No specific announcement at the moment.However, if you are interested in joining the team, please feel free to submit your spontaneous applications to philippe.juin@inserm.fr
F1. 2019-2021 European Commission MSCA-IF A Basseville “Machine learning prediction for breast cancer therapy” - grant ID: 841313- 184 000 €
F2. 2021-2024 French Ligue Against Cancer GO (Regional) A Basseville 3-yr PhD student funding: Development of a new deep learning algorithm (Reductive Discriminating Networks) to improve personalized medicine in breast cancer 94 000 €
F3. 2021-2022 Cancéropôle Grand-Ouest (Regional) PF Cartron Impact of nutrition on epimarks 15 000€
F4. 2021-2026 INCa PrevBIO (National) S Lelièvre (collab. PF Cartron) 226 000€
F5. 2021-2026 ARC Programme Labellisé (National) D Poncet (collab. PF Cartron) 30 000€
F6. 2021-2023 Cancéropôle Grand-Ouest (Regional) PF Cartron/F Souazé Effects of targeted therapies on the stroma 109 000€
F7. 2022 French Ligue Against Cancer GO (Regional) F Souazé “Investigation of therapeutic targets to reverse stromal-mediated, metabolic-adaptation–induced apoptosis resistance in triple-negative breast cancer cells.” 25 000€
F8. 2022 ICO innovation award (Local)- A Basseville - Involvement of Tumor Nerves in Hormone Therapy Response in Breast Cancer Patients: A Spatial Transcriptomics Approach 20 000€
F9. 2022-2024 INCA PLBIO (National) PP Juin Chemotherapeutic response of fibroblast ecosystems in human breast cancer at single cell resolution 191 624 € ( plus 2022 PP Juin Trajectoire nationale Région Pays de la Loire 50 000 €
F10. 2022 SATT Ouest-Valorisation (Regional) PF Cartron Development of epitranscriptomic modulators 420 000€
F11. 2022-2026 ARC SIGN’IT (National) D Fradin (collab PF Cartron), 108 000€
F12. 2023-2027 “Equipe Labellisée” French Ligue Against Cancer (National) PP Juin, “Identification of MCL-1–dependent compensatory mechanisms contributing to cellular heterogeneity in aggressive breast cancers.”120 000 € per year
F13. 2023 Prix de Recherche Ruban Rose (National) PP Juin 35 000 €
F14. 2023- 2027 INCa-DGOS SIRIC ILIAD 2 (National) (led Prof. F Bodere) PP Juin Prog 2 WP6 “Features and vulnerabilities of residual/resisant breast cancer ecotypes” approx 100 000 € per year
F15. 2023-2025 INCA PLBIO (National) N Bidère (collab S Barillé-Nion) - Molecular Basis and cellular Functions of CK1a Interplay with non-canonical IKKs / 36 months 185 328 €
F16. 2025-2027 Odysséa/Persévérance fondation (Local) PF Cartron Long read sequencing 450 000€
F17. 2025 French Ligue Against Cancer GO (Regional) - A Basseville - Involvement of Tumor Nerves in Hormone Therapy Response in Breast Cancer Patients: A Spatial Transcriptomics Approach 30 000€
F18. 2025-2027 INCA PLBIO (National) - M Blondel (collab L Maillet) Role of non-canonical secondary structures of RNA in the alternative splicing of apoptosis regulatory genes and use as intervention points for cancers resistant to chemotherapies. 160 488€
F19. 2025-2027 Cancéropôle Grand-Ouest (Regional) S Barillé-Nion Impact of cell death modalities on therapy induced secretome 15 000€
Facts illustrating the academic appeal or reputation of the team
A1. S Barillé and PF Cartron are co-heads of the Prostate and Breast Cancer and Epigenetics of Tumor Niches Networks of Cancéropôle Grand Ouest (INCa labeled French inter-regional research organization covering three regions, established under the national Plan Cancer) of which PP Juin is Scientific Director
A2. L Maillet is an expert member of CRNS Section 22 (Integrative Structral Biology) since 2025
A3. S Barillé (CSS2), F Souazé (CSS2) and L Maillet (CSS1) were promoted INSERM CHRC in 2022, 2025 and 2026 respectively
A4. V Guen was recruited as INSERM CRCN in 2023 (CSS1) and E Grasset was recruited as CNRS CRCN (Former Section 24) in 2025- see Team DevCan
A5. PP Juin received Ruban Rose Awards in 2012 and 2022 and the "Prix Etancelin de l'Académie des Sciences" in 2021; A Basseville received ICO Innovation Award in 2022
Dr V. Sauzeau, l'Institut du Thorax , Nantes
Dr S Brouard/ A Moreau CR2TI, Nantes
Dr. S. Teletchea US2B, Nantes
Prof F Saubion LERIA, Angers
Prof F Panloup LAREMA, Angers
Drs M Priault/S Manon IBCG, Bordeaux
Prof M Blondel GGB, Brest
Dr C Vandier/Prof O Hérault LNox, Tours
Dr C Ginestier, CRCM, Marseille
Dr. F Mechta-Grigoriou Stress & Cancer , Institut Curie Paris
Dr K Campbell Beatson Institute Glasgow (UK)
Dr SE Bates Columbia University NY (USA)
Visit of the lab
|
||||||||||||||||||||||||||